Markers of steroid receptor, kinase signalling pathways and Ki-67 expression in relation to tamoxifen sensitivity and resistance
Introduction
Resistance to endocrine therapy (ET) is an ongoing clinical challenge in the treatment of estrogen receptor positive ER+ breast cancer (BC). Between 50–80% of ER+ patients initially show clinical benefit to ETs that target estrogen/ER signaling (1,2). Tamoxifen (TAM), a SERM, is one ET that has been used for nearly 40 years in BC (3) and acts by competitively binding to the ER thus blocking the ability of estrogen to drive tumor growth (4). Many patients now receive aromatase inhibitor (AI) treatment, which acts by causing estrogen deprivation, yet TAM has not been superceded by AIs in every clinical scenario, e.g., remaining an important adjuvant ET in premenopausal patients. In postmenopausal patients, 10 years treatment with an AI has not been established as superior to 5 years. Thus, the best extended ET options in post-menopausal BC are still unclear and may incorporate TAM since all of the following have been shown in RCTs to be better than 5 years of TAM alone, i.e., (I) 10 years of TAM (ATLAS & aTTOM) (5,6)], (II) 5 years of TAM followed by 3–5 years of AI (MA17) (7) or (III) 5 years of AI (ATAC, BIG 1–98) (8). TAM still has a significant role in BC chemoprevention (9-12) where studies up to 20 years (9,11) provide data on its long-term efficacy vs. side-effects. It remains clinically-important to identify BCs most likely to be sensitive to TAM.
Steroid hormone receptors, growth factor signaling and resistance to ET
Resistance can either be ‘de-novo’ (i.e., no evidence of hormone sensitivity—defined in the advanced setting as progression within 6 months of starting ET) or ‘acquired’ (i.e., in this setting, after a period of response/stable disease for at least 6 months the tumor progresses). In ER+ BC the proliferation marker Ki-67 is a prognostic tool that may be valuable in defining responsiveness to ET (13,14).
It may be possible to define other molecular predictive biomarkers based on ET signalling. ER-alpha (ER) expression is commonly retained in ET resistance (15) and is mostly in its wild type form in BCs refractory to TAM since its mutation is enriched primarily in estrogen deprivation resistant patients where it is believed to promote a constitutively-active receptor (16-18). A second ER, ER beta, has five isoforms and while the data for ERβ is engaging it has not yet been established as a marker with clinical utility in breast cancer. For example, in one study of 177 postmenopausal women who received adjuvant tamoxifen, the recurrence-free interval was found to differ with respect to degree of nuclear ERβ1 expression (19). In another study from our own research group of 880 patients with primary breast cancer, nuclear ERβ2 and ERβ5, but not ERβ1, significantly correlated with OS and ERβ2 additionally with DFS. ERβ2 also predicted response to endocrine therapy and correlated positively with ERα, progesterone receptor and androgen receptor (20). Not only were the tumor and clinical outcome correlations different between these two studies for ERβ1 but the positivity varied from 33% to 99% respectively, despite the fact both used the same mouse monoclonal antibody clone PPG5/10 which was raised against the unique COOH-terminal peptides of ERβ1. In a further study involving 911 patients with breast cancer, ERβ1 expression (again using this antibody) reported positivity in 89% of tumors (21). In this study, high ERβ1 expression was a favourable prognostic marker, especially in chemotherapy treated patients, but not in the endocrine therapy-treated breast cancer patients. Furthermore, in a pre-surgical study comparing the effect of short-term tamoxifen or three doses of fulvestrant (50, 125 and 250 mg) versus placebo we previously reported significant changes in ERα, PR and Ki-67 (22). However, there was no significant difference between any of the endocrine arms and placebo in terms of either ERβ1 or ERβ2 (personal communication J Robertson). Therefore, while the role of ERβ in breast cancer continues to be explored (23) we did not include it as a biomarker in the present study but focussed on ERα.
Breast cancer models have helped reveal multiple potential mechanisms underpinning resistance (24). Cross talk between ER and deregulated growth factor-driven kinase signaling pathways has been most implicated (25,26), including subverting responsiveness to TAM (27). Such mechanisms have been proposed as a key focus for progress in overcoming endocrine resistance (28). Three membrane receptor tyrosine kinases (RTKs) with reported roles in cell proliferation in particular have been linked to TAM resistance: insulin-like growth factor (IGF) 1 receptor (IGF1R), epidermal growth factor receptor (EGFR), and HER2. Inhibition of such receptors can lead to restoration of the therapeutic effect of TAM (29). IGF1R has been reported in relation to the endocrine responsive phenotype (30). Blockade of the IGFR pathway with the monoclonal antibody Ganitumab showed a negative therapeutic impact (31). Downstream of such RTKs, PI3K signaling through hyper-activation of Akt plays an important role ER+ BC progression, being able to facilitate resistance to ET (32-37). An E17K mutation in AKT1 is also capable of constitutively-activating this enzyme (38). Loss of the enzyme phosphatase and tensin homolog (PTEN) can furthermore result in Akt hyperactivity (39,40) and its loss has been associated with TAM resistance (30). Phosphorylation on Thr246 of Proline-rich Akt substrate of 40 kDA (PRAS40), an Akt substrate and indicator of PI3K pathway activation status (37,41), might also feasibly relate to worsened prognosis but to date there is a paucity of studies exploring the contribution for pPRAS40 in TAM resistant ER+ BC.
Several routes exist for subsequent cross-talk between deregulated RTK/kinase pathways and ER, including ligand independent activation of ER (24). Nuclear ER phosphorylation of AF-1 residues including serine 118 and 167 (pER118 and pER167 respectively) can be triggered by these kinase pathways. Non-genomic (plasma membrane) ER interacts with RTKs activating downstream kinases that can drive such ligand-independent nuclear ER phosphorylation. ERK1/2 MAP kinase (MAPK) has been linked to induced phosphorylation of ER’s serine 118 residue, thereby promoting hormone independent growth in models (42,43). Hyperactive Akt can phosphorylate ER on serine 167, for example when driven by PTEN loss (40). Ligand-independent ER activation in turn may enhance expression of growth factors (44,45). Such signaling can be causal in BC models resistant to ET (28) including TAM (44,45).
An alternative nuclear steroid hormone receptor, the androgen receptor (AR), a relatively new marker in relation to BC response to ET, is potentially important (46). The expression of AR has been related to expression of ER (47) and subsequently in a larger series of BCs associated with multiple phenotypes and biomarkers (48). AR has also been reported as a prognostic marker (49). Thus, AR positive tumors are associated with luminal A disease (50), low grade tumors (50,51), and better outcome in patients with ER+ BC (49,50). In contrast, AR overexpression has been linked with TAM resistance in pre-clinical experiments (52,53); Cochrane et al. reported that the risk of TAM failure for patients with a nuclear AR/ER ratio ≥2 was 4 times that of the lower ratio subgroup (54).
To identify if any of these molecular markers implicated in steroid receptor and RTK/kinase pathways (including PI3K/Akt signaling) predicts response or resistance to TAM in patients with ER+ BC 10 biomarkers were immunohistochemically-evaluated. By including biopsy specimens and tumor microarrays (TMAs), as well as from different tumor locations, the study was able consider whether biomarker data derived from different intra-tumoral locations or sample types can be combined for such studies. We present the following article in accordance with the MDAR checklist (available at http://dx.doi.org/10.21037/tbcr-20-31).
Methods
Patients and immunohistochemical
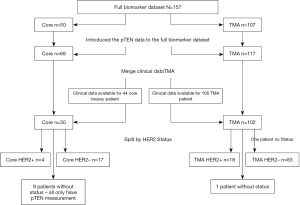
A total of 123 endocrine treatment naïve patients with known ER and HER2 status were treated with TAM and had clinical outcome data: 102 had their tumor samples in a tumor microarray format (TMAs) and 21 as core biopsies (Figure 1)]. An additional five patients with TMAs but no outcome data were included only in the analysis of biomarker expression in TMA samples from different intra-tumoral locations (i.e., central and peripheral) (i.e., total TMA samples n=107).
Patients reported here were commenced on TAM between 1983–2002. During the earlier part of this period patients with ER unknown tumors or even on occasion known ER negative indolent disease were given TAM. For this report, in keeping with the ASCO guidelines, ER+ has been defined as ≥1% of tumor cells staining positive (55). 103/123 (84%) patients had ER+ tumors and of these 83/103 patients had surgery as initial treatment of their tumors. For these 83 patients, three 0.6 mm TMA samples were prepared from two central and one peripheral regions from the resected tumor tissue block for each patient: these patients received no adjuvant ET and were treated with TAM when they developed recurrent, advanced BC. The remaining 20 ER+ patients who did not have initial surgery for their tumors had core biopsies only, and were treated with TAM as a primary ET: these were either elderly patients with primary BC and were being treated by primary ET or patients who presented with advanced BC for whom TAM was the initial ET. Survival analyses included only ER+ patients with TAM clinical outcome data (n=103) (Figure 1). Categorization by HER2 status is also shown in Figure 1. Clinical data included measurements for time to progression (TTP), time to death (TTD), duration of clinical benefit (DoCB), clinical benefit (CB) and objective response (OR).
All tissue evaluated was formalin-fixed paraffin-embedded (FFPE) tumor material and specimens and clinical data were provided by the University of Nottingham and City Hospital (now Queens University Hospital), Nottingham. The study was conducted in accordance with the Declaration of Helsinki (revised 2013) and the Harmonized Tripartite Guideline for Good Clinical Practice from the International Conference on Harmonization. Informed consent was waived and archival tissues were used with approval of the Ethics Committee and R & D Committee. The research protocol had approved from Nottingham Research Ethics Committee (Ref: 2020313) and R & D approval from the hospital (Ref: 03H101).
Biomarker measurements were performed by the University of Nottingham, the Breast Cancer Molecular Pharmacology Group in Cardiff University, and the Division of Oncology and Pathology, Department of Clinical Sciences in Lund University, Sweden. Ten biomarkers were evaluated by immunohistochemical (IHC) staining (assessing % positivity and/or H-Score), reflecting steroid receptor and RTK-kinase signaling as well as proliferation, comprising: ER-alpha (ER), HER2, Ki-67, phosphorylated (p) forms of ERK1/2 MAPK, Akt, ER (Ser118), PRAS40, and IGF1R, and PTEN & AR expression. All assays were applied to the clinical BC TMAs and core biopsies, except PTEN which was measured on core biopsies and full tumor sections (FTS) and AR which was measured only on TMAs (Table 1). Table S1 details the monoclonal antibodies used, phosphorylation site detected and IHC. A minimum of 100 tumor epithelial cells were required for assessment of the IHC marker staining. Eight IHC markers were evaluated in Cardiff University using antibodies for ER expression, pER118, pMAPK, Ki-67, HER2, pIGF1R, pPRAS40 and pAkt followed by assessment by consensus of 2 experienced personnel (JMW Gee, P Finlay) blinded to the clinical data. All assays were performed by running small batches of test slides, together with an in-house internal assay positive breast cancer control to monitor for adequate IHC assay staining performance in each assay and so ensure no marked inter-assay differences. Each TMA was run in separate assays along with a subset of the biopsy samples. PTEN staining was performed in Nottingham. For this, staining was assessed by one observer and an H-score measured. Equivocal or borderline cases were re-examined and a consensus score was reached with a second observer. AR staining was performed and assessed in Lund by one observer. PTEN and AR IHC methodologies are again shown in Table 1. Table 1 shows the cellular location of staining (n = nuclear, c = cytoplasmic, m = plasma membrane) evaluated and the scoring method for all biomarkers. Figure S1 shows examples of higher and weaker staining for the various markers.
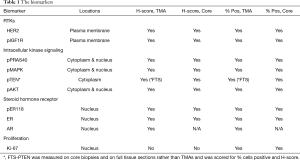
Full table
Each tumor was also analysed for Akt1 E17K somatic mutation using BEAMing (Beads, Emulsions, Amplification, and Magnetics) technology by Sysmex Inostics GmbH, Hamburg, Germany (56).
Analysis cohorts
The analysis cohorts consisted of (I) ER+ patients only, (II) ER+ patients sub-grouped by HER2 status (i.e., luminal A: ER+/HER2− & luminal B ER+/HER2+) and iii) grouped according to tissue type (core biopsy or TMA; also TMA combined with core biopsy) (Figure 1). HER2m status was categorized into negative “0” ≤30% or positive “1” >30% according to % of 3+ complete membrane staining. For each patient, the clinical TAM outcome & levels of staining for each biomarker comprised the analysis data.
Given the size of the dataset it was necessary to combine the IHC scores from the TMA and core biopsies for analysis versus outcome for each marker. Previous publications, including from our own unit, have reported that core biopsies and surgical excision specimens show good correlation for ER status (57-59) with the H-score only slightly lower (between 10–20 on a 0–300 scale) in the surgical specimen compared to the core biopsy (57,58). A meta-analysis of 21 published studies involving almost 2,500 patients also reported good overall concordance between core and surgical excision of 92.8% for ER and 85.2% for PR (60). Fewer prior publications exist regarding biomarkers of RTK-kinase pathways (61), in particular phospho-specific markers compared according to type of sample, although one publication has reported some differences (62).
In exploring biomarker relationship to TAM outcome it was thus first important to be aware of any impact of combining steroid receptor, RTK/kinase or proliferation data from the different types of sample in the present study. This was achieved by evaluating differences in staining for individual biomarkers between TMAs and core biopsies (which in this study was an inter-patient, inter-tumor comparison). To more robustly achieve this, the number of core biopsies for staining was further increased by including further diagnostic pre-treatment core biopsies that were available without clinical outcome data (thus giving a total number of cores for this analysis n=50). The results were evaluated by non-parametric testing (Wilcoxon test applies throughout) before false discovery rate (FDR) correction in an exploratory analysis of sample positivity rate and staining level within the TMAs versus core biopsies for the biomarkers. Exploratory correlations were evaluated between levels of the signaling biomarkers to gauge whether the biological observations noted were also rational for the combined TMA plus core biopsy sample set.
Biomarkers in TMAs (n=107) obtained from the two central and the peripheral regions of the tumors were similarly statistically-examined, this time to evaluate any biomarker differences according to sample location within a tumor (i.e., intra-patient, intra-tumoral comparison) and so gauge if sample location might influence biomarker findings. The two central and one peripheral value were subsequently combined to get a single TMA value for each patient by taking the mean of all three values (central a, central b, peripheral) for analysis versus outcome.
Statistical analysis of IHC biological (steroid receptor, RTK-kinase and proliferation) endpoints versus patient outcome
The ability of the IHC biomarkers to predict response to TAM as measured by CB, TTP, and TTD was examined in the ER+ patient cohort and according to HER2 status or sample type. In addition to analyzing the full set of IHC markers, 3 further analyses were performed examining a combination of ER & Ki-67, pAkt and pPRAS40 (as an indicator of active Akt signaling) and also analysis of AR/ER ratio.
The potential role of the markers in prediction of response to TAM was explored through logistic and survival models of clinical outcomes. Univariate and multivariate methods were used to model the markers individually and in combination. Logistic regression and Cox proportional hazards models were used to model clinical benefit and survival endpoints respectively. FDR correction was applied to results from univariate analyses, and variable selection methods including regression trees were used to avoid over fitting the multivariate model. Bootstrap methods (resampling from the original data) and model selection based on AIC were used to validate variable selection for logistic models and the final model for survival analyses. Interquartile range (IQR) odds ratios and IQR hazard ratios are presented along with confidence intervals, illustrating difference between high and low (3rd and 1st quartiles) measures for each of the markers.
In the absence of any predefined cut points, the biomarkers were considered continuous predictors. Significance was set at one sided with P<0.05.
Results
Exploratory analyses of biological (steroid receptor, RTK/kinase & proliferation) IHC endpoints
Analysis of the IHC biomarker positivity rate in core biopsy vs. TMA samples
Table 2 includes the number (and %) of biopsy samples (core or TMA)—showing any degree of immuno-positivity for each marker (considering n, m or c staining localization). As stated above, 50 core biopsies and up to 107 TMAs (with or without clinical outcome data) were included in this exploratory analysis. pIGFRm had substantially more positive core biopsies than positive TMAs and for pAKTn, pER118n, pMAPKc, pMAPKn and pPRAS40c somewhat increased positivity rate was also seen in core biopsies. No marked discordance was noted for the % tumors showing any immunopositivity for total nuclear ER (ERn), HER2m, pAktc or Ki-67 between core biopsy vs. TMA sample types. Only pPRAS40n had a reduced % positivity rate in the core biopsies compared with TMA samples (Table 2). PTEN was measured on either FTS or core biopsies and therefore was not included in this comparison of TMAs versus core biopsies. AR was not measured in cores, but showed a substantial % nuclear positivity rate in the TMAs (Table 2).
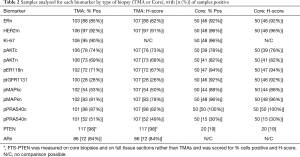

Full table
Analysis of IHC biomarker staining level in core biopsy vs. TMA samples and inter-marker correlations in the combined dataset
The positive staining level (H-Score or %) in the TMAs compared with core biopsies was different for some of the biomarkers; when FDR correction was applied to the resultant individual Wilcoxon P values the hypothesis that % positivity or H-scores cannot be assumed to be the same was verified (Table 3). Staining values were generally lower in TMAs, and after FDR correction significant differences remained for pER118n, pIGF1Rm, pMAPKc, HER2m, pAktc and pPRAS40c, and for ER H-Score only, but there were no significant differences for pAktn, pMAPKn or pPRAS40n, or for ER or Ki-67 when measured by percentage positivity, between core biopsy and TMA samples.
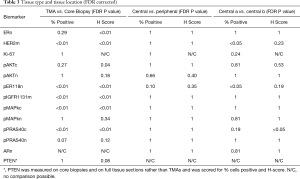
Full table
Exploratory analysis revealed several biologically-expected significant correlations between markers in the combined biopsy cores and TMAs (Figure S2), notably between total and pER118n; for pMAPKn, pAktn and pPRAS40c with pER118n; between pAktc and pPRAS40c, and several for pIGF1Rm (including with HER2m, PRAS40c and pER118n). An inverse correlation was noted between ER expression and Ki-67 (Figure S2).
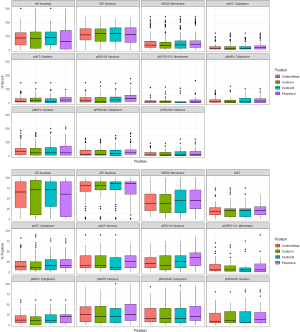
Comparing the IHC biomarkers in TMAs obtained from central & peripheral regions of tumors
Figure 2 show the box plots of % positivity and H-score respectively comparing each of the different sample locations within each tumor for the biomarkers. After FDR correction there was no significant difference between the peripheral sample and the mean of the two central TMA samples for % positivity or H-Score of any of the markers (Table 3). Between the two central samples, there was a modest significant difference (P<0.05) detected for % positivity only for HER2m and pER118n (Table 3) while for H-score there was a modest significant difference (P<0.05) detected for pPRAS40c (Table 3). The lack of marked differences support feasibility of combining staining data for these biomarkers (including Ki-67) at the different tumor sites (as a mean) for analysis versus outcome
Analyses of Akt1 E17K somatic mutation
Only 2/123 (1.6%) tumors had Akt1 E17K mutations.
Relationships between IHC biomarkers and tamoxifen outcome
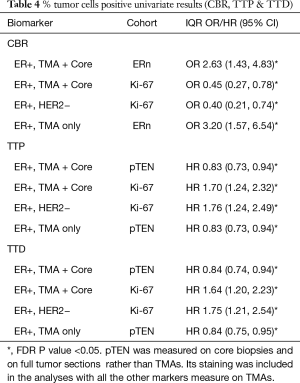
While the IHC biomarkers were scored by % positive staining and H-Score, the majority of the significant results versus outcome pertained to the % positive results (Table 4). There were also too few patients in the HER2+ cohort to obtain conclusive data, so the following results refer only to the full ER+ combined TMA + Core biopsy cohort, the ER+ TMA only cohort, and the luminal A (ER+/HER2−, TMA + Core biopsy) cohort.
Full table
Clinical benefit on tamoxifen
In the ER+ combined TMA + Core biopsy sample cohort, univariate models suggested there was an increased odds of clinical benefit (CB) associated with decreased levels of Ki-67 and increased levels of ERn expression, as measured by % positivity (Table 4). Given the differences in some biomarkers (other than Ki-67) between core biopsy and TMAs, studies were also performed within the TMA cohort alone, where the observation with ERn was retained. The significant association with Ki-67 was also seen in the luminal A (ER+/HER2−) cohort using the combined (ER+ TMA+ core biopsy) cohort. After adjustment for FDR no other significances for any of the RTK-kinase signaling molecules were obtained versus CB (Table 4). To avoid overfitting in the multivariate model, variable selection methods (regression trees of 1,000 bootstrapped samples) were used; the only markers included in the multivariate model were ERn and Ki-67, and also pPRAS40c and pAkt (as an indicator of active Akt signaling). Only Ki-67 and ER % positivity were retained as significant for CBR in the final multivariate model for the full sample cohort (Table 5). In the Luminal A cohort only the Ki-67 association remained when analyzing the full ER+ sample set (Table 4).
Full table
While AR alone was not predictive, the AR/ER staining ratio revealed 61 had a ratio <2, and 10 had a ratio ≥2. Fishers exact test showed that the odds of achieving CB was enriched in patients with an AR/ER ratio <2 [P<0.05, odds ratio 5.1 (1.01, 33.4)].
Survival endpoints following tamoxifen treatment
Univariate models were also fitted for TTP and TTD. After FDR correction a significant increased hazard of disease progression (TTP) and death (TTD) was seen in the full ER+ and also the Luminal A cohorts for those with higher Ki-67. Within the full ER+ cohort (FTS+ core biopsy or FTS only, but not for the luminal A cohort) a significantly increased risk of disease progression and death was found for those with lower PTEN % positivity levels (Table 4). No other associations were seen for the survival endpoints.
Multivariate models analyses were performed using the full ER+ cohort, considering the Core biopsy + TMA combined or TMA alone samples, although as noted above pTEN was measured in FTS and core biopsies: in both analyses increased Ki-67 and decreased PTEN were predictive of TTP and TTD (Table 5). Only Ki-67 proved significant versus these endpoints in the Luminal A cohort. In contrast to the CB data, no significant results were seen for TTP and TTD in relation to AR/ER ratio. These results were validated by bootstrapping and model selection based on AIC.
Discussion
This study adds to the literature on the impact of type of clinical sample and of intra-tumoral site of biopsy when assessing biomarkers as predictors of clinical utility in breast cancer. Even after FDR correction significant differences in staining level remained for the majority of markers according to sample type (pER118n, pIGF1Rm, pMAPKc, HER2m, pAktc and pPRAS40c and for ER H-Score only). There were no significant differences for pAktn, pMAPKn, pPRAS40n, Ki-67 or ER when assessed by percentage positivity or H-score between core biopsy and TMAs. This is an important and reassuring finding since ER and Ki-67 are two of the markers identified as significantly associated with clinical outcome in this study and associations with outcome have also been noted for Ki-67 in sequential sample studies (13). Differences commonly noted in signaling marker staining for immunopositivity rate and level between core biopsies and TMAs (often staining higher in the former) may be related to differences in fixation/preparation of each type of sample influencing antigenicity and thus the subsequent IHC assay data. This has previously been suggested by other groups (61,62), who reported somewhat higher levels of selected AKT signaling elements and phospho-MAPK in core biopsies. It should however be appreciated that in the present study the core biopsy versus excision specimens were not a direct comparison from the same tumor and so differences reported here may, at least in part, also be due to differences in phenotype of tumors which had core biopsy (primary ET patients) versus TMAs constructed from surgically resected tumors. Importantly, however, there was no significant difference in Ki-67 between TMA and core samples.
For all the biomarkers reported in the present study the findings suggest that the site biopsied within a tumor is not a critical issue: our analysis of multiple biomarkers was a direct comparison within the same tumor. This is important in interpreting the results of studies using percutaneous core biopsy material where differences in biomarkers between groups have been suggested may be due, at least in part, to the sampling of different sites within the tumors. Equally studies assessing the effect of drug therapies using sequential core biopsies in an individual (22,63) have reported differences in expression/activity of the various steroid receptor and RTK/kinases. Our study would indicate that such differences are not due to sequential biopsies being taken from different sites within the tumor. These findings are in keeping with the study by Meric-Bernstam and colleagues who reported no significant difference in 7 biomarkers measured by IHC according to tumor location (62): Ki-67, PTEN & p4EBP1 T70 showed strong correlations between central and peripheral expression, pAktS473, pPRAS40 T246 and pS6 S240/244 showed moderate correlations, and only pS6 S235/236 showed a low correlation. This study also measured biomarkers by RPPA and reported that of 154 markers measured, 132 (86%) were not statistically different between the tumor centre and periphery. Tramm and colleagues recently reported on 27 patients assessing five biomarkers in central and peripheral regions, each within two separate tumor blocks (64): there was almost perfect concordance for BCL2, E-cadherin and EGFR between the central and peripheral regions and moderate/substantial agreement for EMMPRIN and Ki-67. Nevertheless, it should be noted that the finding of similar biomarker expression in peripheral and central sites in a tumor remains a matter of debate. Douglas-Jones and colleagues reported differences in ER expression across the width of tumors (58). Likewise, Aleskandarany and colleagues reported heterogeneity of Ki-67 expression within a tumor (although this was between ‘full face’ sections from different primary tumor blocks rather than from central or peripheral TMA cores within the tumor) (65). There may also be some loss of antigen signal between core biopsies and surgical resected specimens due to technical factors, particularly for phosphorylated proteins (62): where antigen signal has been diminished this might provide some explanation for similarity in signal between different areas of the same resected tumor specimen.
The above highlight the importance of describing the study methodology and technical aspects of biomarker assessment when trying to correlate biomarker expression with clinical utility. We found using univariate and multivariate analysis that measures of clinical response to TAM can be predicted by increased staining for ER, the PI3K/Akt pathway regulator PTEN, or lower Ki-67, either in combination or individually. This was seen in the full ER+ cohort. For the ER+ TMA + Core combined group PTEN, Ki-67 and ERn as measured by % positive staining were all predictive: ERn & Ki-67 predicted CBR whereas Ki-67 and PTEN predicted TTP and TTD. It should be noted that ER and Ki-67 were also two markers for which there was no evidence of difference between TMA and core biopsy samples (in terms of % positivity) or between central or peripheral sites of the tumor, indicating these may be a robust combination of predictive markers for TAM outcome irrespective of sample type. PTEN was measured on FTS and core biopsies, so there was no central versus peripheral TMA comparison for PTEN in our study. However, Meric-Bernstam and colleagues (62) reported “strong correlation” between PTEN expression of central and peripheral samples, suggesting this may also be a reliable signaling marker. Within the luminal A cohort only decreased Ki-67 was seen to be predictive of increased CB, TTP, and TTD while increased pTEN was predictive of an increased TTP & TTD.
Our ER, PTEN and Ki-67 predictive biomarker findings were not influenced by AKT mutation status, since such mutation was very low in this BC (1.6%), a finding reflective of the literature since a positivity rate for AktE17K mutation frequencies of 6.3% has been reported (66), and where previous published literature ranges from 1.4–8.2%, with a mean mutation frequency of 3.8% (67).
With regards to AR/ER ratio, the patient numbers were small and so the relationship detected between the <2 ratio and clinical benefit of TAM should be viewed with caution. Nevertheless, since they are in agreement with studies from Cochrane and colleagues where AR/ER ratio correlates with clinical outcomes (54), our findings suggest further research would be worthwhile to confirm or refute whether there is a meaningful relationship between AR/ER ratio and TAM clinical outcome.
Several biologically-plausible correlations were preserved between the signaling steroid receptor and RTK-kinase biomarkers in this TMA plus core biopsy series. Of note, the correlations between total ER and pER118n, and between pMAPKn and pER118n, are in keeping with ER signaling and kinase/ER cross talk via ER activation (42,43). Correlations were also seen between pAkt and pPRAS40, elements both lying in the PI3K/Akt signaling pathway. Furthermore, the inverse correlation noted between ER and Ki-67 has been previously observed (68). However, despite these reassuring biological correlations and the wealth of supportive preclinical data from ET resistant BC models, unfortunately other than PTEN we found none of the examined markers reflective of RTK-kinase signaling and its cross-talk with ER were related to TAM outcome clinically. On one hand this might be surprising since inhibition of the PI3KCA/Akt/mTOR pathway has been reported to result in significant increases in TTP/PFS (69,70). In these other clinical studies, most of the patients had been heavily pre-treated. Equally there are studies where inhibitors of these pathways have shown no significant improvement in TTP/PFS (71,72). The fact that the present study included hormone naïve patients and also used primary tumor tissue, where the PI3KCA/Akt/mTOR pathway may not be as disrupted as in metastases (73-75), may explain why no further markers reflective of the Akt pathway were intrinsically associated with TAM resistance. This therefore is one limitation of this study and it again highlights the importance of obtaining tumor tissue immediately prior to a new treatment especially within future studies. Likewise, studies comparing the molecular phenotype of primary and metastases within the same patient are important to elucidate the issues related to using primary tumors to predict response at a much later time point. A recent meta-analysis of 5,000 patients with HR+ advanced BC showed that liver visceral metastases have a worse outcome than other sites of metastases (76,77) suggesting it may also be important to biopsy metastases from different organ sites to best predict clinical outcome to targeted therapies.
It would be difficult to biopsy all sites of metastases in routine clinical practice but from a perspective of trying to understand mechanism of resistance this could be viewed as another potential limitation. Furthermore, biopsies taken prior to starting any treatment, whether from the original primary tumor or metastases, may allow prediction of response or ‘de-novo’ progression. However, to more fully understand the mechanisms of acquired resistance, which can occur over months to many years, multiple sequential tumor biopsies on treatment and at relapse must be studied (78-81). The present study therefore was not able to look for markers which would highlight potential mechanisms of acquired resistance.
From a mechanistic perspective, the observation that EGFR/HER2/IGF1R signalling and its potential cross-talk via kinases with ER phosphorylation were not biomarkers of tamoxifen resistance in our study might also be explained by the abundance of further resistance mechanisms that have emerged from laboratory studies. For example, along with many changes in ER signalling (including changes in ER’s potential dimer partners, ER variants and ER coactivators such as AIB1 (82,83), the de-regulation of alternative kinase signalling pathways and transcription factors [e.g., FGFR (84,85)], changes in non-coding RNAs (86,87), cell cycle regulators and apoptotic machinery (88) are just some of the mechanisms that have been proposed to contribute to endocrine resistance. The establishment and use of breast cancer cell lines that are tamoxifen resistant in vitro has been valuable towards the discovery and verification of many resistance mechanisms, with some of these models also having substantial proliferation in the presence of tamoxifen (89) and in some instances reduced ER (90) and PTEN loss (29,91) as observed in the present clinical study. Indeed, several of the deregulated pathways, following promising pre-clinical studies, have progressed to being evaluated and (in the case of CDK4/6 inhibition (92) approval as therapeutic targets alongside endocrine agents. Studies of patient tumor DNA samples and more complex in vivo model systems are also linking endocrine resistance to tumor heterogeneity, underpinned by further mechanisms involving genetic/epigenetic events (93), tumor microenvironment and cancer stem cells (94,95). Some of these mechanisms are likely to be important contributors to the control of proliferative activity and might also influence ER level, limiting effectiveness of agents such as tamoxifen.
Conclusions
In conclusion, while we acknowledge the limitation of this analysis due to the available sample size our results in this study support previous work that high ER and low Ki-67 are predictive of clinical utility in hormone naïve patients receiving TAM and also adds PTEN loss. Significantly, considering the findings in this study together with other studies of biopsy type and site (62), all three markers appear independent of sample type and tumor region, adding to the robustness of these markers in relation to TAM outcome. In contrast, for the remaining 7 markers, it is feasible that some of these markers might have had some value if measured within a more advanced phenotype, since type of sample and the disease setting may be critical in relation to prediction of ET response/ progression. It may be more informative for biomarkers to be assessed on biopsies taken just prior to a particular therapy, with analytical variables minimized, in order to best gauge prediction of response in that setting. This should be our aim in prospective randomized trials of new targeted treatments in advanced BC. Furthermore, it remains important that we continue to uncover potential new targets and relevant markers from the most promising of the mechanisms of resistance emerging in experimental studies. In the future, these should continue to be prioritized for evaluation alongside ER, Ki-67 and PTEN as potential biomarkers to aid in selecting patients most likely to derive durable tamoxifen benefit.
Acknowledgments
The authors acknowledge funding support from Bayer AG for this study which was performed as an investigator-initiated study. Acknowledgement is also made to Drs Andrea Haegebarth and Oliver Politz of Bayer AG for scientific discussions during the project.
Funding: None.
Footnote
Reporting Checklist: The authors have completed the MDAR reporting checklist. Available at http://dx.doi.org/10.21037/tbcr-20-31
Data Sharing Statement: Available at http://dx.doi.org/10.21037/tbcr-20-31
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at http://dx.doi.org/10.21037/tbcr-20-31). JFR serves as an unpaid editorial board member of Translational Breast Cancer Research from Feb 2020 to Jan 2022. PF accepted the funded from grant by Bayer AG; JMWG’s lab work was funded by grant from Bayer AG; JFRR was projected funded by grant from Bayer AG. The authors have no other conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. The study was conducted in accordance with the Declaration of Helsinki (revised 2013) and the Harmonized Tripartite Guideline for Good Clinical Practice from the International Conference on Harmonization. Informed consent was waived and archival tissues were used with approval of the Ethics Committee and R & D Committee. The research protocol had approved from Nottingham Research Ethics Committee (Ref: 2020313) and R & D approval from the hospital (Ref: 03H101).
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Goldhirsch A, Colleoni M, Gelber RD. Endocrine therapy of breast cancer. Ann Oncol 2002;13:61-8. [Crossref] [PubMed]
- Robertson JFR, Llombart-Cussac A, Rolski J, et al. Activity of fulvestrant 500 mg versus anastrozole 1 mg as first-line treatment for advanced breast cancer: results from the FIRST study. J Clin Oncol 2009;27:4530-5. [Crossref] [PubMed]
- Osborne CK, Schiff R. Mechanisms of endocrine resistance in breast cancer. Annu Rev Med 2011;62:233-47. [Crossref] [PubMed]
- Tate AC, Greene GL, DeSombre ER, et al. Differences between estrogen and antiestrogen receptor complexes from human breast tumors identified with an antibody raised against the estrogen receptor. Cancer Res 1984;44:1012-8. [PubMed]
- Davies C, Pan H, Godwin J, et al. Long-term effects of continuing adjuvant tamoxifen to 10 years versus stopping at 5 years after diagnosis of oestrogen receptor-positive breast cancer: ATLAS, a randomised trial. Lancet 2013;381:805-16. [Crossref] [PubMed]
- Gray RG, Rea D, Handley K, et al. ATTom: Long-term effects of continuing adjuvant tamoxifen to 10 years versus stopping at 5 years in 6,953 women with early breast cancer. J Clin Oncol 2013;31:abstr 5.
- Goss PE, Ingle JN, Martino S, et al. A randomized trial of letrozole in postmenopausal women after 5 years of tamoxifen therapy for early-stage breast cancer. N Engl J Med 2003;349:1793-802. [Crossref] [PubMed]
- Early Breast Cancer Trialists' Collaborative Group (EBCTCG). Aromatase inhibitors versus tamoxifen in early breast cancer: patient-level meta-analysis of the randomised trials. Lancet 2015;386:1341-52. [Crossref] [PubMed]
- Powles TJ, Ashley S, Tidy A, et al. Twenty-year follow up of the Royal Marsden randomized, double-blinded tamoxifen breast cancer prevention trial. J Natl Cancer Inst 2007;99:283-90. [Crossref] [PubMed]
- Fisher B, Costantino JP, Wickerham DL, et al. Tamoxifen for the prevention of breast cancer: current status of the National Surgical Adjuvant Breast and Bowel Project P-1 study. J Natl Cancer Inst 2005;97:1652-62. [Crossref] [PubMed]
- Cuzick J, Stetak I, Cawthorn S, et al. Tamoxifen for prevention of breast cancer: extended long-term follow-up of the IBIS-1 breast cancer prevention trial. Lancet 2015;16:67-75. [Crossref] [PubMed]
- Veronesi U, Maisonneuve P, Rotmensz N. Italian randomized trial among women with hysterectomy: tamoxifen and hormone-dependent breast cancer in high-risk women. J Natl Cancer Inst 2003;95:160-65. [Crossref] [PubMed]
- Dowsett M, Smith IE, Ebbs SR, et al. Prognostic value of Ki-67 expression after short term presurgical endocrine therapy for primary breast cancer. JNCI 2007;99:167-70. [Crossref] [PubMed]
- Weigel MT, Dowsett M. Current and emerging biomarkers in breast cancer: prognosis and prediction. Endocr Relat Cancer 2010;17:R245-62. [Crossref] [PubMed]
- Robertson JFR. Oestrogen receptor: a stable phenotype in breast cancer. Br J Cancer 1996;73:5-12. [Crossref] [PubMed]
- Jeselsohn R, Yelensky R, Buchwalter G, et al. Emergence of constitutively active estrogen receptor-a mutations in pretreated advanced estrogen receptor-positive breast cancer. Clin Cancer Res 2014;20:1757-67. [Crossref] [PubMed]
- Carlson KE, Choi I, Gee A, et al. Altered ligand binding properties and enhanced stability of a constitutively active estrogen receptor: evidence that an open pocket conformation is required for ligand interaction. Biochemistry 1997;36:14897-905. [Crossref] [PubMed]
- Lazennec G, Ediger TR, Petz LN, et al. Mechanistic aspects of estrogen receptor activation probed with constitutively active estrogen receptors: correlations with DNA and coregulator interactions and receptor conformational changes. Mol Endocrinol 1997;11:1375-86. [Crossref] [PubMed]
- Reese JM, Suman VJ, Subramaniam M, et al. ERβ1: characterization, prognosis, and evaluation of treatment strategies in ERα-positive and -negative breast cancer. BMC Cancer 2014;14:749. [Crossref] [PubMed]
- Shaaban AM, Green AR, Karthik S, et al. Nuclear and cytoplasmic expression of ERbeta1, ERbeta2, and ERbeta5 identifies distinct prognostic outcome for breast cancer patients. Clin Cancer Res 2008;14:5228-35. [Crossref] [PubMed]
- Elebro K, Borgquist S, Rosendahl AH, et al. High Estrogen Receptor beta Expression Is Prognostic among Adjuvant Chemotherapy- Treated Patients—Results from a Population-Based Breast Cancer Cohort. Clin Cancer Res 2017;23:766-77. [Crossref] [PubMed]
- Robertson JFR, Nicholson RI, Bundred NJ, et al. Comparison of the short-term biological effects of 7alpha-[9-(4,4,5,5,5-pentafluoropentylsulfinyl)-nonyl]estra-1,3,5,(10)-triene-3,17beta-diol (Faslodex) versus tamoxifen in postmenopausal women with primary breast cancer Cancer Res 2001;61:6739-46. [PubMed]
- Omoto Y, Iwase H. Clinical significance of estrogen receptorβ in breast and prostate cancer from biological aspects. Cancer Sci 2015;106:337-43. [Crossref] [PubMed]
- Dixon JM. Endocrine resistance in breast cancer. New J Sci 2014. [Crossref]
- Nicholson RI, McClelland RA, Robertson JFR, et al. Involvement of steroid hormone and growth factor cross-talk in endocrine response in breast cancer. Endocr Relat Cancer 1999;6:373-87. [Crossref] [PubMed]
- Schiff R, Massarweh SA, Shou J, et al. Cross-talk between estrogen receptor and growth factor pathways as a molecular target for overcoming endocrine resistance. Clin Cancer Res 2004;10:331S-6S. [Crossref] [PubMed]
- Osborne CK, Shou J, Massarweh S, et al. Crosstalk between Estrogen Receptor and Growth Factor Receptor Pathways as a cause for Endocrine Therapy Resistance in Breast Cancer. Clin Cancer Res 2005;11:865S-70S. [PubMed]
- García-Becerra R, Santos N, Diaz L, et al. Mechanisms of resistance to endocrine therapy in breast cancer: Focus on signalling pathways, miRNAs and genetically based resistance. Int J Mol Sci 2012;14:108-45. [Crossref] [PubMed]
- Miller TW, Perez-Torres M, Narasanna A, et al. Loss of Phosphatase and tensin homologue deleted on chromosome 10 engages ErB3 and IGF-1R signalling to promote antiestrogen resistance in breast cancer. Cancer Res 2009;69:4192-201. [Crossref] [PubMed]
- Zhang Y, Moerkens M, Ramaiahgari S, et al. Elevated insulin-like growth factor 1 receptor signaling induced antiestrogen resistance through the MAPK/ERK and PI3K/Akt signaling routes. Breast Cancer Res 2011;13:R52. [Crossref] [PubMed]
- Robertson JFR, Ferrero JM, Bourgeois H, et al. Ganitumab with either exemestane or fulvestrant for postmenopausal women with advanced, hormone receptor-positive breast cancer: a randomised, controlled, double-blind, phase 2 trial. Lancet Oncol 2013;14:228-35. [Crossref] [PubMed]
- Campbell RA, Bhat-Nashatri P, Patel NM, et al. Phosphatidylinositol 3-kinase/Akt-mediated activation of estrogen receptor. Journal of Biological Chemistry 2001;276:9817-24. [Crossref] [PubMed]
- Pérez-Tenorio G, Stal O. Activation of Akt/PKB in breast cancer predicts a worse outcome among endocrine treated patients. Br J Cancer 2002;86:540-5. [Crossref] [PubMed]
- Osaki M, Oshimura M, Ito H. PI3K-Akt pathway: its functions and alterations in human cancer. Apoptosis 2004;9:667-76. [Crossref] [PubMed]
- Tokunaga E, Kimura Y, Mashino K, et al. Activation of PI3K/Akt signaling and hormone resistance in breast cancer. Breast Cancer 2006;13:137-44. [Crossref] [PubMed]
- Ghayad SE, Vendrell JA, Ben Larbi S, et al. Endocrine resistance associated with activated ErbB system in breast cancer cells is reversed by inhibiting MAPK or PI3K/Akt signalling pathways. Int J Cancer 2010;126:545-62. [Crossref] [PubMed]
- Wang H, Zhang Q, Wen Q, et al. Proline-rich Akt substrate of 40 kDa (PRAS40): A novel downstream target of PI3K/Akt signaling pathway. Cell Signal 2012;24:2226. [Crossref]
- Carpten JD, Faber AL, Horn C, et al. A transforming mutation in the pleckstrin homology domain of AKT1 in cancer. Nature 2007;448:439-44. [Crossref] [PubMed]
- Carracedo A, Pandolfi PP. The PTEN-PI3K pathway: of feedbacks and cross-talks. Oncogene 2008;27:5527-41. [Crossref] [PubMed]
- Vilgelm A, Lian Z, Wang H, et al. AKtmediated phosphorylation and activation of estrogen receptor is required for endometrial neoplastic transformation in pTEN +/− Mice. Cancer Res 2006;66:3375-80. [Crossref] [PubMed]
- Manning BD, Cantley LC. Akt/PKB signaling: navigating downstream. Cell 2007;129:1261-74. [Crossref] [PubMed]
- Kato S, Endoh H, Masuhiro Y, et al. Activation of the estrogen receptor through phosphorylation by mitogen-activated protein kinase. Science 1995;270:1491-4. [Crossref] [PubMed]
- Kurokawa H, Lenferink AEG, Simpson JF, et al. Inhibition of HER2/ neu (erb B-2) and Mitogen-activated protein kinases enhances tamoxifen action against HER2-overexpressing, tamoxifen-resistant breast cancer cells. Cancer Research 2000;60:5887-94. [PubMed]
- Britton DJ, Hutcheson IR, Knowlden JM, et al. Bidirectional cross talk between ERalpha and EGFR signalling pathways regulates tamoxifen-resistant growth. Breast Cancer Res Treat 2006;96:131-46. [Crossref] [PubMed]
- Knowlden JM, Hutcheson IR, Barrow D, et al. Insulin-like growth factor-I receptor signaling in tamoxifen-resistant breast cancer: a supporting role to the epidermal growth factor receptor. Endocrinology 2005;146:4609-18. [Crossref] [PubMed]
- Fioretti FM, Sita-Lumsden A, Bevan CL, et al. Revising the role of the androgen receptor in breast cancer. J Mol Endocrinol 2014;52:R257-65. [Crossref] [PubMed]
- Isola JJ. Immunohistochemical demonstration or androgen receptor in breast cancer and its relationship to other prognostic factors. J Pathol 1993;170:31-5. [Crossref] [PubMed]
- Collins L C, Kimberly SC, Marotti JD, et al. Androgen receptor expression in breast cancer in relation to molecular phenotype: results from the Nurses’ Health Study. Mod Pathol 2011;24:924-931. [Crossref] [PubMed]
- Vera-Badillo FE, Templeton AJ, de Gouveia P, et al. Androgen receptor expression and outcomes in early breast cancer: a systematic review and meta-analysis. JNCI 2014;106:djt319. [Crossref] [PubMed]
- Park S, Koo JS, Kim MS, et al. Androgen receptor expression is significantly associated with better outcomes in estrogen receptor-positive breast cancers. Ann Oncol 2011;22:1755-62. [Crossref] [PubMed]
- Park S, Koo J, Park HS, et al. Expression of androgen receptors in primary breast cancer. Ann Oncol 2010;21:488-492. [Crossref] [PubMed]
- Ciupek A, Rechoum Y, Gu G, et al. Androgen receptor promotes tamoxifen agonist activity by activation of EGFR in ER_-positive breast cancer. Breast Cancer Res Treat 2015;154:225-37. [Crossref] [PubMed]
- De Amicis F, Thirugnansampanthan J, Cui Y, et al. Androgen Receptor Overexpression Induces Tamoxifen Resistance in Human Breast Cancer Cells. Breast Cancer Res Treat 2010;121:1-11. [Crossref] [PubMed]
- Cochrane DR, Beranles S, Jacobsen BM, et al. Role of the androgen receptor in breast cancer and preclinical analysis of enzalutamide. Breast Cancer Res 2014;16:R7. [Crossref] [PubMed]
- Hammond ME, Hayes DF, Dowsett M, et al. American Society of Clinical Oncology/College of American Pathologists Guideline Recommendations for Immunohistochemical Testing of Estrogen and Progesterone Receptors in Breast Cancer (Unabridged Version). Arch Pathol Lab Med 2010;134:e48-72. [PubMed]
- Dressman D, Yan H, Traverso G, et al. Proceedings of the National Academy of Sciences USA 2013;100:8817-22. [Crossref]
- Hodi Z, Chakrabarti J, Lee AHS, et al. The reliability of assessment of oestrogen receptor expression on needle core biopsy specimens of invasive carcinomas of the breast. J Clin Pathol 2007;60:299-302. [Crossref] [PubMed]
- Douglas-Jones AG, Collett N, Morgan JM, et al. Comparison of core oestrogen receptor (ER) assay with excised tumour: intratumoral distribution of ER in breast carcinoma. J Clin Pathol 2001;54:951-5. [Crossref] [PubMed]
- Zidan A, Brown JSC, Peston D, et al. Oestrogen and progesterone receptor assessment in core biopsy specimens of breast carcinoma. J Clin Pathol 1997;50:27-9. [Crossref] [PubMed]
- Li S, Yang X, Zhang Y, et al. Assessment accuracy of core needle biopsy for hormone receptors in breast cancer: a meta-analysis. Breast Cancer Res Treat 2012;135:325-34. [Crossref] [PubMed]
- Pinhel IF, Macneill FA, Hills MJ, et al. Extreme loss of immunoreactive p-Akt and p-Erk1/2 during routine fixation of primary breast cancer. Breast Cancer Res 2010;12:R76. [Crossref] [PubMed]
- Meric-Bernstam F, Akcakanat A, Chen H, et al. Influence of biospecimen variables on proteomic biomarkers in breast cancer. Clin Cancer Res 2014;20:3870-83. [Crossref] [PubMed]
- Robertson JFR, Coleman RE, Cheung KL, et al. AZD5363, an AKT inhibitor, significantly inhibits key biomarkers of the AKT pathway and Ki-67 in a randomized, placebo controlled study (STAKT) in human breast cancers. Clin Cancer Res 2020;26:1574-85. [Crossref] [PubMed]
- Tramm T, Kyndi M, Sørensen FB, et al. Influence of intra-tumoral heterogeneity on the evaluation of BCL2, E-cadherin, EGFR, EMMPRIN, and Ki-67 expression in tissue microarrays from breast cancer. Acta Oncol 2018;57:102-6. [Crossref] [PubMed]
- Aleskandarany MA, Green AR, Ashankyty I, et al. Impact of intratumoural heterogeneity on the assessment of Ki-67 expression in breast cancer. Breast Cancer Res Treat 2016;158:287-95. [Crossref] [PubMed]
- Rudolph M, Anzeneder T, Schulz A, et al. AKT1E17K mutation profiling in breast cancer: prevalence, concurrent oncogenic alterations, and blood-based detection. BMC Cancer 2016;16:622. [Crossref] [PubMed]
- Troxell ML. PIK3CA/AKT1 mutations in breast carcinoma: a comprehensive review of experimental and clinical studies. J Clin Exp Pathol 2012;S1:002.
- Ragab HM, Samy N, Afify M, et al. Assessment of Ki-67 as a potential biomarker in patients with breast cancer. J Genet Eng Biotechnol 2018;16:479-84. [Crossref] [PubMed]
- Yardley DA, Noguchi S, Pritchard KI, et al. Everolimus Plus Exemestane in Postmenopausal Patients with HR+ Breast Cancer: BOLERO-2 Final Progression-Free Survival Analysis. Adv Ther 2013;30:870-84. [Crossref] [PubMed]
- André F, Ciruelos EM, Rubovszky G, et al. Alpelisib (ALP) + fulvestrant (FUL) for advanced breast cancer (ABC): Results of the phase III SOLAR-1 trial. Ann Oncol 2018;29:mdy424.010.
- Wolff AC, Lazar AA, Bondarenko I, et al. Randomized Phase III Placebo-Controlled Trial of Letrozole Plus Oral Temsirolimus As First-Line Endocrine Therapy in Postmenopausal Women With Locally Advanced or Metastatic Breast Cancer. J Clin Oncol 2013;31:195-202. [Crossref] [PubMed]
- Baselga J, Dent SF, Cortés J, et al. Phase III study of taselisib (GDC-0032) + fulvestrant (FULV) v FULV in patients (pts) with estrogen receptor (ER)-positive, PIK3CA-mutant (MUT), locally advanced or metastatic breast cancer (MBC): Primary analysis from SANDPIPER. J Clin Oncol 2018. [Crossref]
- Beelen K, Hoefnagel LD, Opdam M, et al. PI3K/AKT/mTOR pathway activation in primary and corresponding metastatic breast tumors after adjuvant endocrine therapy. Int J Cancer 2014;135:1257-63. [Crossref] [PubMed]
- Schleifman EB, Desai R, Spoerke JM, et al. Targeted biomarker profiling of matched primary and metastatic estrogen receptor positive breast cancers. PLoS One 2014;9:e88401. [Crossref] [PubMed]
- Dupont Jensen J, Laenkholm AV, Knoop A, et al. PIK3CA mutations may be discordant between primary and corresponding metastatic disease in breast cancer. Clin Cancer Res 2011;17:667-77. [Crossref] [PubMed]
- Robertson JFR, Paridaens R, Bogaerts J, et al. Meta-analyses of visceral versus non-visceral metastases treated by SERM, AI & SERD agents as 1st line endocrine therapy (ET) for HR+ breast cancer (BC). Cancer Res 2019;15:P4-13-11.
- Robertson JFR, Di Leo A, Johnston S, et al. Meta-analyses of visceral versus non-visceral metastases treated by AI & SERD agents as 2nd line endocrine therapy (ET) for HR+ breast cancer (BC). Cancer Res 2019. [Crossref]
- Kenny FS, Pinder S, Ellis IO, et al. Gamma linolenic acid with tamoxifen as primary therapy in breast cancer. Int J Cancer 2000;85:643-8. [Crossref] [PubMed]
- Kenny FS, Willsher PC, Gee JMW, et al. Change in expression of ER, bcl-2 and MIB1 on primary tamoxifen and relation to response in ER positive breast cancer. Breast Cancer Res Treat 2001;65:135-44. [Crossref] [PubMed]
- Robertson JFR, Gutteridge E, Cheung KL, et al. Oestrogen reception expression in human breast cancer during long-term fulvestrant treatment. J Clin Oncol 2004;22:536. [Crossref]
- Agrawal A, Robertson JF, Cheung KL, et al. Biological effects of fulvestrant on estrogen receptor positive human breast cancer: short, medium and long-term effects based on sequential biopsies. Int J Cancer 2016;138:146-59. [Crossref] [PubMed]
- Girault I, Bieche I, Lidereau R. Role of estrogen receptor alpha transcriptional coregulators in tamoxifen resistance in breast cancer. Maturitas 2006;54:342-51. [Crossref] [PubMed]
- Haque MM, Desai KV. Pathways to Endocrine Therapy Resistance in Breast Cancer. Front Endocrinol (Lausanne) 2019;10:573. [Crossref] [PubMed]
- Turner N, Pearson A, Sharpe R, et al. FGFR1 amplification drives endocrine therapy resistance and is a therapeutic target in breast cancer. Cancer Res 2010;70:2085-94. [Crossref] [PubMed]
- Zhou Y, Yau C, Gray JW, et al. Enhanced NF kappa B and AP-1 transcriptional activity associated with antiestrogen resistant breast cancer. BMC Cancer 2007;7:59. [Crossref] [PubMed]
- Miller TE, Ghoshal K, Ramaswamy B, et al. MicroRNA-221/222 confers tamoxifen resistance in breast cancer by targeting p27kip1. J. Biol. Chem 2008;283:29897-903. [Crossref] [PubMed]
- Farhan M, Aatif M, Dandawate P, et al. Non-coding RNAs as Mediators of Tamoxifen Resistance in Breast Cancers. Adv Exp Med Biol 2019;1152:229-41. [Crossref] [PubMed]
- Thangavel C, Dean JL, Ertel A, et al. Therapeutically activating RB: reestablishing cell cycle control in endocrine therapy-resistant breast cancer. Endocr Relat Cancer 2011;18:333-45. [Crossref] [PubMed]
- Knowlden JM, Hutcheson IR, Jones HE, et al. Elevated levels of epidermal growth factor receptor/c-erbB2 heterodimers mediate an autocrine growth regulatory pathway in tamoxifen-resistant MCF-7 cells. Endocrinology 2003;144:1032-44. [Crossref] [PubMed]
- Stone A, Valdés-Mora F, Gee JM. PLoS One 2012;7:e40466. [Crossref] [PubMed]
- Phuong NT, Kim SK, Lim SC, et al. Role of PTEN promoter methylation in tamoxifen-resistant breast cancer cells. Breast Cancer Res Treat 2011;130:73-83. [Crossref] [PubMed]
- Cristofanilli M, Turner NC, Bondarenko I, et al. Fulvestrant plus palbociclib versus fulvestrant plus placebo for treatment of hormone-receptor-positive, HER2-negative metastatic breast cancer that progressed on previous endocrine therapy (PALOMA-3): final analysis of the multicentre, double-blind, phase 3 randomised controlled trial. Lancet Oncol 2016;17:425-39. [Crossref] [PubMed]
- Razavi P, Chang MT, Xu G, et al. The Genomic Landscape of Endocrine-Resistant Advanced Breast Cancers. Cancer Cell 2018;34:427-38.e6. [Crossref] [PubMed]
- Piggott L, Silva A, Robinson T, et al. Acquired Resistance of ER-Positive Breast Cancer to Endocrine Treatment Confers an Adaptive Sensitivity to TRAIL through Posttranslational Downregulation of c-FLIP. Clin Cancer Res 2018;24:2452-63. [Crossref] [PubMed]
- Jallow F, O'Leary KA, Rugowski DE, et al. Dynamic interactions between the extracellular matrix and estrogen activity in progression of ER+ breast cancer. Oncogene 2019;38:6913-25. [Crossref] [PubMed]
Cite this article as: Campbell C, Mathew J, Ellis IO, Bradbury I, Borgquist S, Elebro K, Green AR, Finlay P, Gee JMW, Robertson JFR. Markers of steroid receptor, kinase signalling pathways and Ki-67 expression in relation to tamoxifen sensitivity and resistance. Transl Breast Cancer Res 2020;1:29.